Diet and gut microbiome interactions of relevance for symptoms in irritable bowel syndrome
We published a study entitled “Diet and gut microbiome interactions of relevance for symptoms in irritable bowel syndrome” in Microbiome journal.
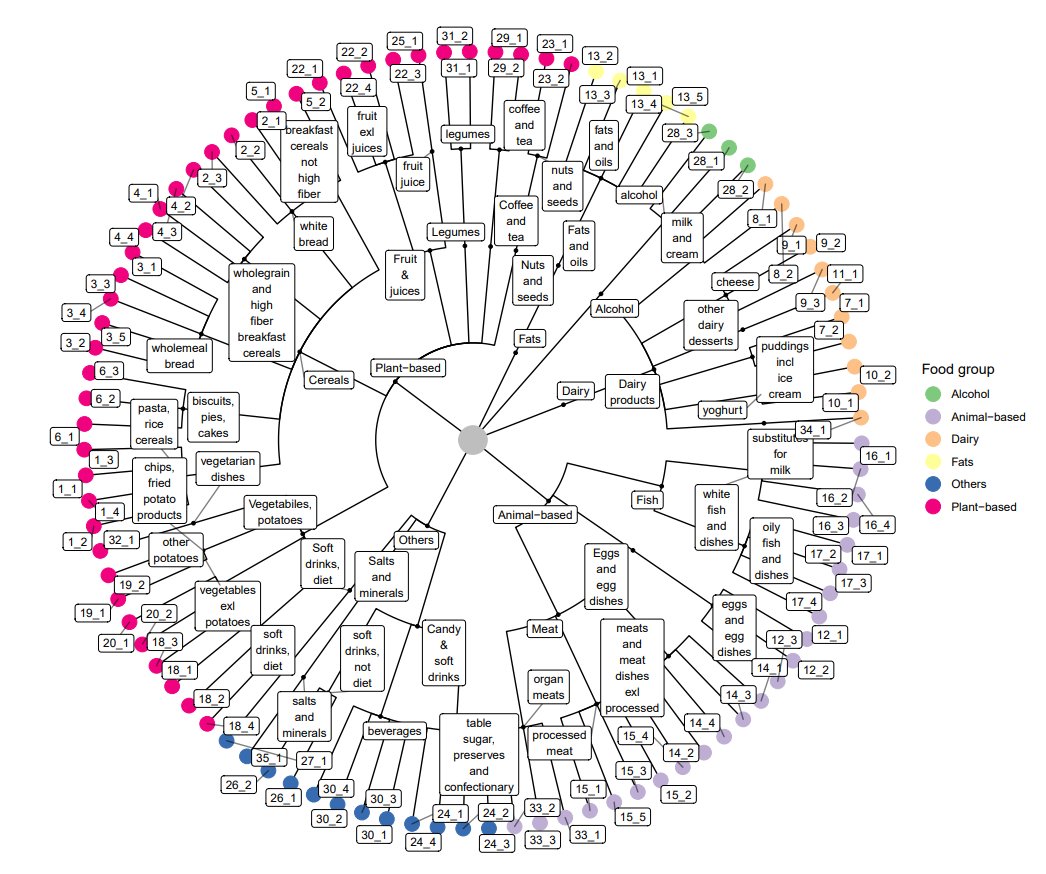
First, we used a food tree approach1 to explore dietary data from our cohort. We added a specific nutrient layer to this food tree and computed FSA-NPS diet index (used now to compute NutriScore in front pack food in Europe).
We notably observed that individuals with severe IBS symptoms consumed a higher amount of lower-quality food items during their main meals than healthy controls.
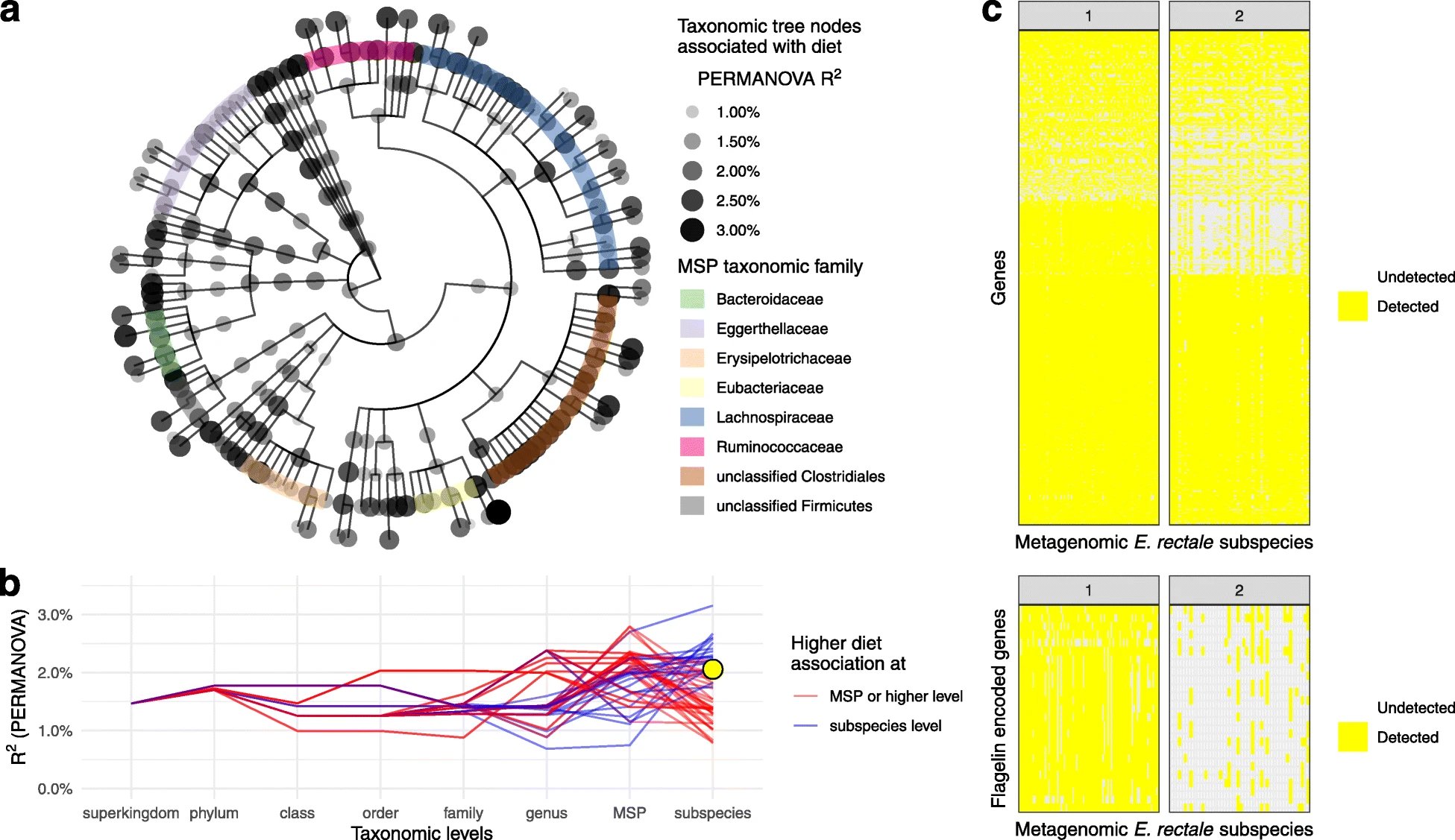
Thank to shotgun metagenomics combining with MSP pangenomes2, we were able to study diet interactions at subspecies levels. We found an interesting signal with Eubacterium rectale flagelin and diet for instance.
Using co-inertia analysis between unifrac food tree and MSP at subspecies levels combining with machine learning, we were able to predict meat/plant ratio, exhaled gas and symptoms severity.
As we found some signals in exhaled gas, carbohydrate metabolism and IBS severity, we aimed to do a specific focus on gut microbial hydrogenase and found a differential signal that correlate with meat/mucin associated CAZy.
Our study3 provides an unprecedented resolution of diet-microbiota-symptom interactions and ultimately guides new interventional studies that aim to identify gut microbiome-based nutritional recommendations for the management of gastrointestinal symptoms.
Johnson AJ et al. Daily sampling reveals personalized diet-microbiome associations in humans. 2019. Cell Host Microbe ↩
Plaza Oñate F et al. MSPminer: abundance-based reconstitution of microbial pan-genomes from shotgun metagenomic data.. 2019. Bioinformatics ↩
Tap J et al. Diet and gut microbiome interactions of relevance for symptoms in irritable bowel syndrome.2021. Microbiome ↩