
Julien is currently a research scientist at INRAE Micalis institute settled in Paris-Saclay campus, where he works on projects related to the interactions between food and human microbiomes in a nutrition and health perspective.

Julien is currently a research scientist at INRAE Micalis institute settled in Paris-Saclay campus, where he works on projects related to the interactions between food and human microbiomes in a nutrition and health perspective.
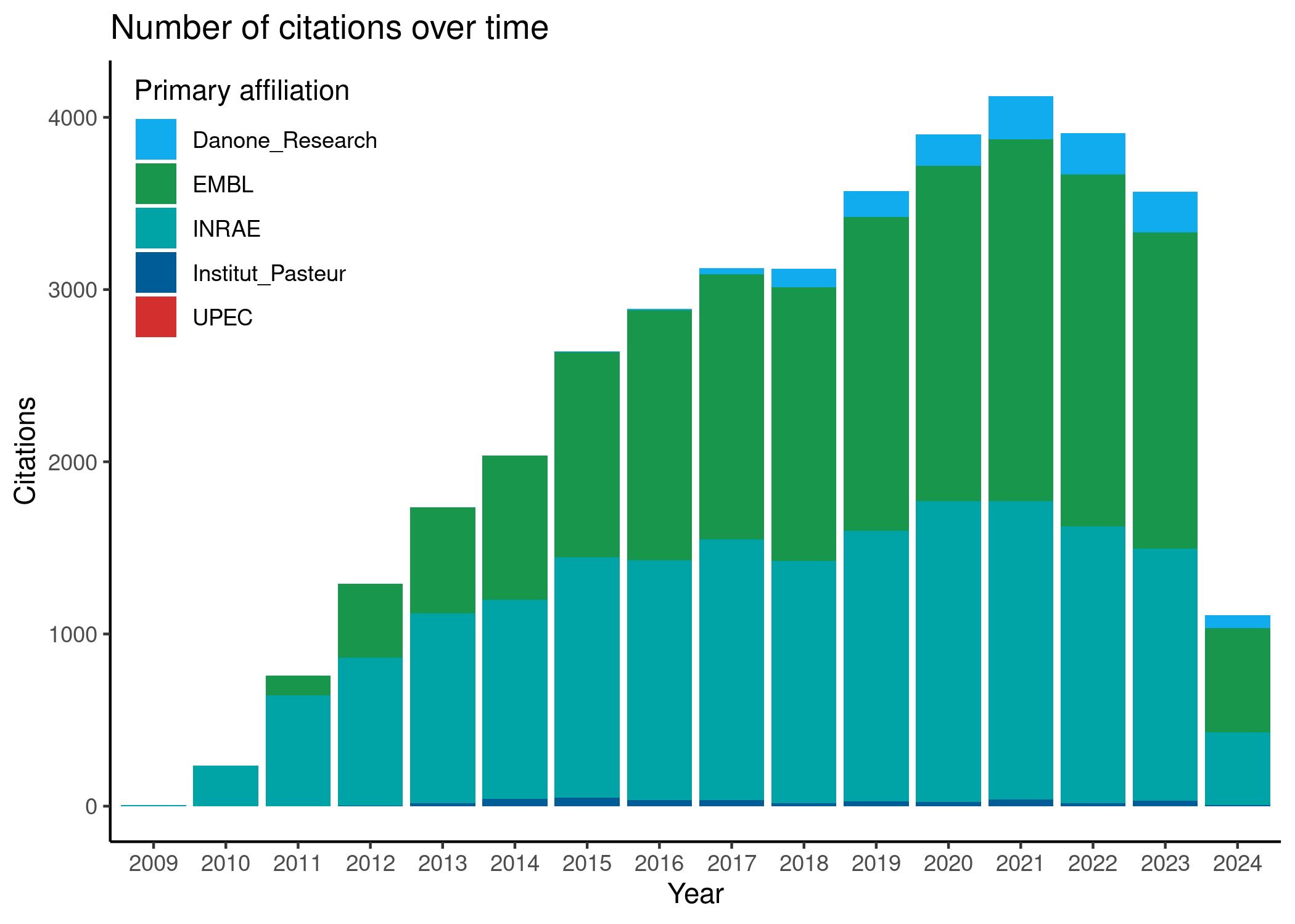
Total Citations: 42729.0
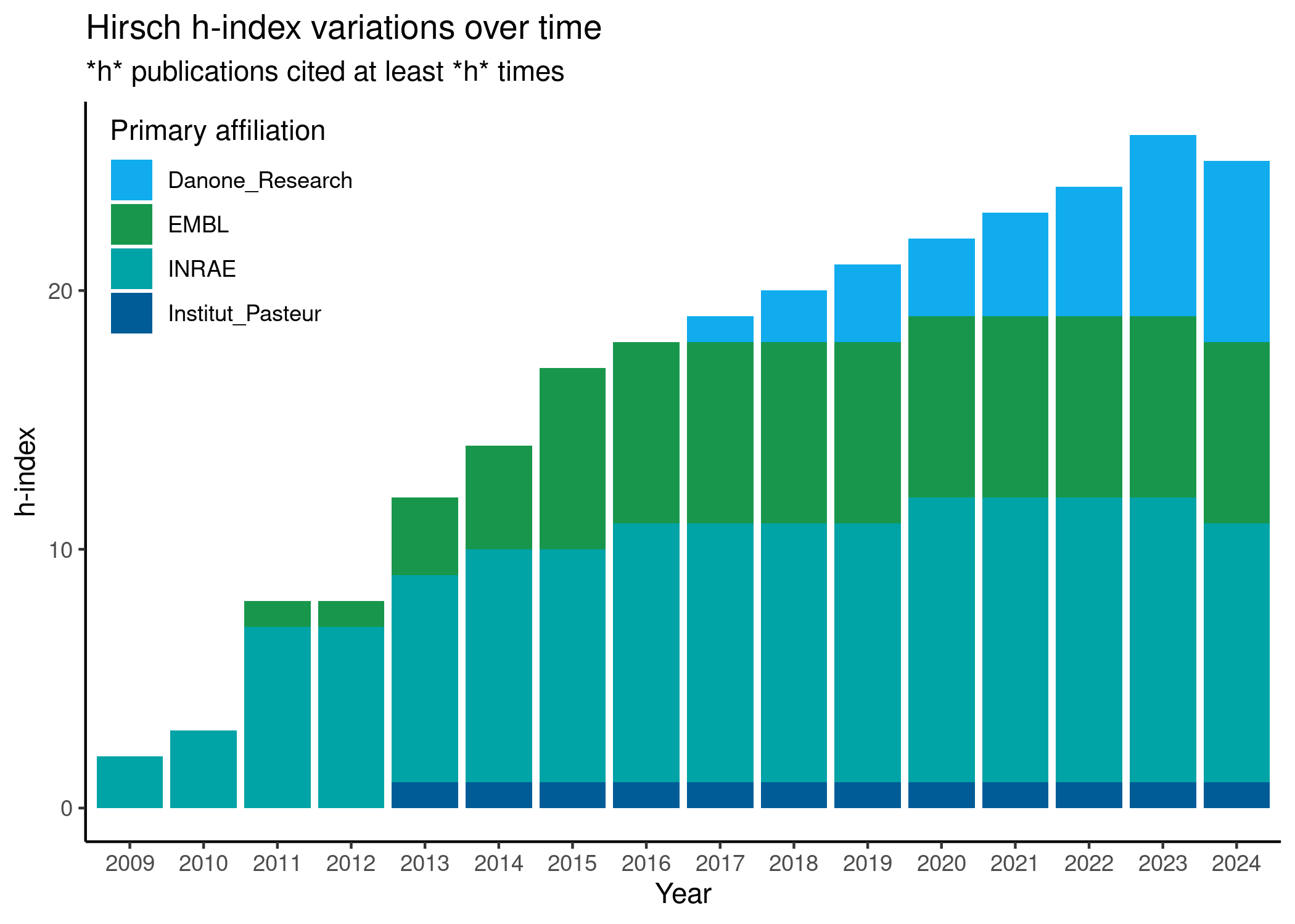
h-index: 28.0
Selected publications