Genomic variation landscape of the human gut microbiome
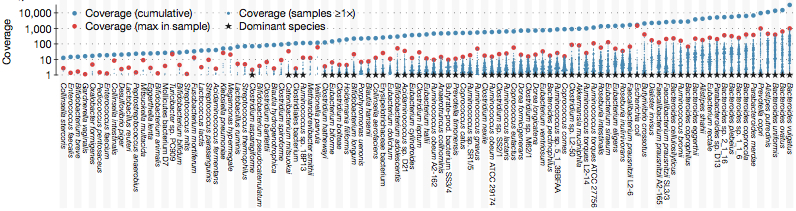
A joint effort between EMBL and WashU scientist have permitted to make genomic analysis on metagenomic data using some ecological metadata (geography, time sampling) ...
Published on December 14, 2012 by Julien Tap
species human gut variation individual genomic
1 min READ

A joint effort between EMBL and WashU scientist have permitted to make genomic analysis on metagenomic data using some ecological metadata (geography, time sampling). This new frame work permit to open new kinds of analyses on gut microbiota. Using 207 public available metagenomic dataset, it had been possible to describe the whole SNP (single nucleotide polymorphism) variation of abundant bacterial species present in the human gut. Two main findings were highlighted :
- Each individual have the same selection pressure on their microbes while each species evolved differently : this permit notably to describe particular gene evolving differently regading different species.
- Each individual have its own microbial stable genotype : while species abundance could vary, it appears that each individual have their own genotype by species…. this observation reinforced the intimate co-evolution between human host and his symbiotic microbiota
Genomic variation landscape of the human gut microbiome. Siegfried Schloissnig, Manimozhiyan Arumugam, Shinichi Sunagawa, Makedonka Mitreva, Julien Tap, Ana Zhu, Alison Waller, Daniel R. Mende, Jens Roat Kultima, John Martin, Karthik Kota, Shamil R. Sunyaev, George M. Weinstock and Peer Bork